New Method Unveils RNA Structural Diversity, Enhancing Understanding of Gene Regulation in Viruses and Fungi
April 24, 2026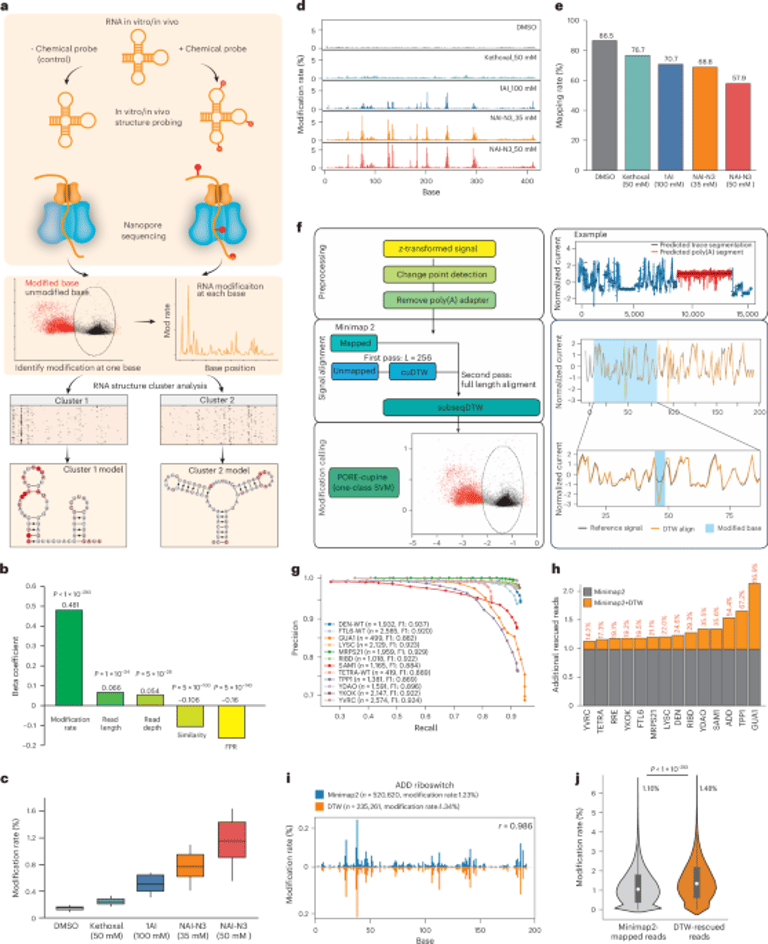
A new single-molecule RNA structure probing method, sm-PORE-cupine, accurately separates RNA structure ensembles for SARS-CoV-2 sgRNAs and the Candida albicans transcriptome, revealing substantial structural heterogeneity that depends on cellular context.
sm-PORE-cupine integrates SHAPE chemical probing with direct nanopore RNA sequencing to read modification patterns on individual RNA molecules, enabling identification of RNA structure ensembles both in cells and in solution.
In SARS-CoV-2, regions rich in RNA–RNA interaction hubs show greater structural heterogeneity, whereas highly homogeneous regions have fewer hubs, with the 3′ end of the genome being particularly heterogeneous due to sgRNA diversity.
In Candida albicans, RNA structure heterogeneity correlates with GC content and translation efficiency, and 3′ UTR heterogeneity decreases at higher temperatures; heat shock drives a global shift toward more homogeneous structures, while specific 3′ UTR regions modulate translation as shown by luciferase assays.
Across transcriptomes, RNA structure ensembles are highly heterogeneous, with differing heterogeneity in 3′ UTRs and coding sequences; translation changes and temperature shifts (heat shock) influence structure homogeneity and regulatory outcomes.
Clustering per-molecule reactivity profiles, notably with a Bernoulli mixture model, reveals distinct structural populations in riboswitches (ADD, YKOK, GUA1) and can distinguish ligand-bound from ligand-free states with high accuracy.
The study supports that RNA structural ensembles dynamically regulate gene expression in eukaryotes, and that single-molecule structure profiling can reveal these dynamics at the scale of the transcriptome.
The analysis uses dynamic time warping–based signal alignment and one-class SVM to map modified RNA signals, increasing read recovery and mappability, with strong concordance to traditional reactivity measurements.
Summary based on 1 source