Breakthrough in Protein Misfolding: New Class Linked to Alzheimer's and Parkinson's Discovered
August 9, 2025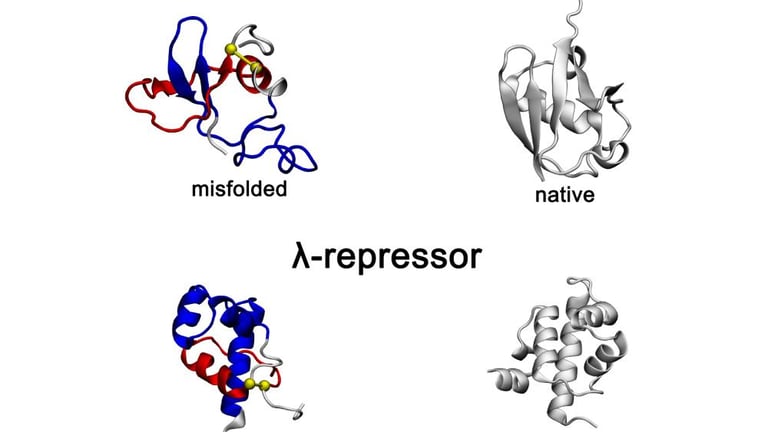
Proteins, composed of amino acids, must fold into specific three-dimensional shapes to function correctly, and any misfolding can disrupt this essential process.
In contrast, larger proteins may harbor these persistent non-native entanglements, which require extensive backtracking to rectify, potentially impairing cellular function.
Researchers at Penn State University have made a significant breakthrough in understanding protein misfolding, identifying a new class termed non-native entanglement, which is linked to diseases like Alzheimer's and Parkinson's.
The research team combined simulations with experimental methods like mass spectrometry, confirming the presence of predicted entangled misfolds and aligning experimental observations with simulation results.
Led by Ed O’Brien, the study aims to elucidate the mechanisms of protein misfolding to identify potential therapeutic targets for related disorders, with findings published in the journal Science Advances.
O'Brien emphasized the importance of comprehending these misfolding mechanisms for developing therapies that could mitigate the impacts of aging and related diseases.
This discovery opens new avenues for research into the molecular mechanisms behind protein aggregation diseases, potentially leading to novel therapeutic strategies.
The study highlights the significance of integrating computational and experimental methods in understanding complex biological processes, marking a notable advancement in protein science.
Non-native entanglement occurs when segments of a protein's amino acid chain become improperly looped or threaded, which can lead to structural changes that evade cellular quality control systems.
The study revealed that while misfolds in smaller proteins are typically short-lived, normal-sized proteins exhibit persistent misfolds due to their hidden nature and the complexity involved in correcting such entanglements.
The findings suggest that existing protein quality control systems are less effective at detecting these topological misfolds, which may contribute to protein accumulation and cellular stress.
Employing advanced all-atom molecular dynamics simulations, the research captured detailed atomic interactions that previous coarse-grained models could not, enhancing the understanding of protein folding.
Summary based on 3 sources
Get a daily email with more Science stories
Sources

Penn State University • Aug 8, 2025
New class of protein misfolding simulated in high definition | Penn State University
News-Medical • Aug 9, 2025
All-atom models reveal novel protein misfolding mechanism
BIOENGINEER.ORG • Aug 8, 2025
High-Definition Simulations Reveal New Class of Protein Misfolding