Breakthrough Study Unveils Dynamic RNA Self-Assembly, Paving Way for AI-Driven Drug Discovery
November 27, 2025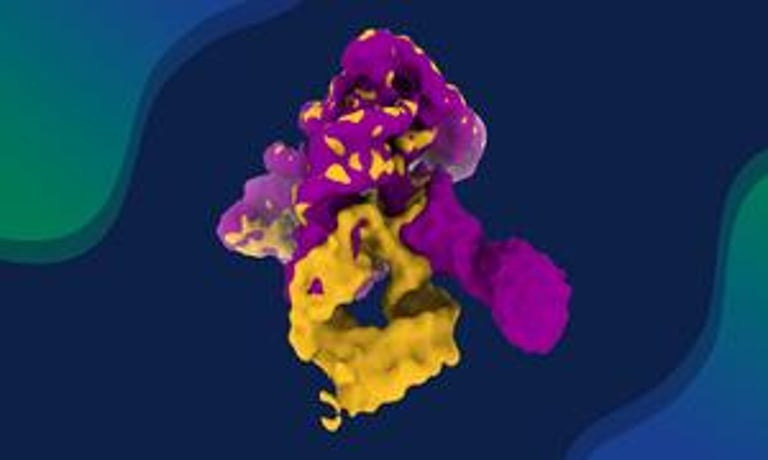
A Nature Communications study offers the most complete view yet of how a self-splicing ribozyme folds and assembles, using an integrative structural biology approach to visualize intermediate states as it becomes catalytically active.
The team combines cryo-EM, SAXS, RNA biochemistry and enzymology, image processing, and molecular simulations to capture the ribozyme’s progressive self-assembly while avoiding kinetic traps.
Experts say the work clarifies dynamic assembly at atomic detail and could influence RNA design, therapeutics, and nanotechnology, potentially accelerating RNA-based drug discovery.
The study lays groundwork for AI-driven RNA structure prediction, offering datasets and benchmarks that could inform future algorithms akin to an AlphaFold for RNA.
By integrating experimental data with machine learning, the research could enable an AI-based approach to RNA structure prediction and model validation.
SAXS data and molecular dynamics simulations reveal low energy barriers between conformations, supporting smooth transitions and refining each structural frame for dynamic modeling.
The combination of SAXS and MD simulations highlights conformational plasticity and enables accurate modeling of the ribozyme’s dynamic transitions.
Overall, the study delivers the most complete molecular film of RNA self-assembly to date and provides benchmarks for AI modeling and RNA engineering.
The research connects the evolution of RNA-based life, noting Group II introns as ancestors of the spliceosome, with practical implications for RNA design, therapeutics, and nanotechnology.
Domain 1 acts as the central scaffold, coordinating the sequential joining of Domains 2, 3, and 4 to prevent misfolding and ensure correct assembly of the catalytic ribozyme.
Collaborations with EMBL Grenoble, CSSB Hamburg, and IIT provided advanced facilities and expertise in cryo-EM, image processing, and molecular simulations, enabling a high-resolution view of the assembly process.
The study analyzes hundreds of thousands of RNA molecules to reconstruct intermediate conformations, enabled by novel cryo-EM image-processing strategies that reveal fleeting frames static structures miss.
Summary based on 3 sources
Get a daily email with more Science stories
Sources

Phys.org • Nov 27, 2025
RNA in action: Filming ribozyme self-assembly
EurekAlert! • Nov 27, 2025
RNA in action: Filming ribozyme self-assembly
News-Medical • Nov 27, 2025
Researchers capture a ribozyme in motion for the first time