TECHNO Platform Revolutionizes Gene Humanization with High-Efficiency CRISPR and BAC-Based Editing
January 17, 2026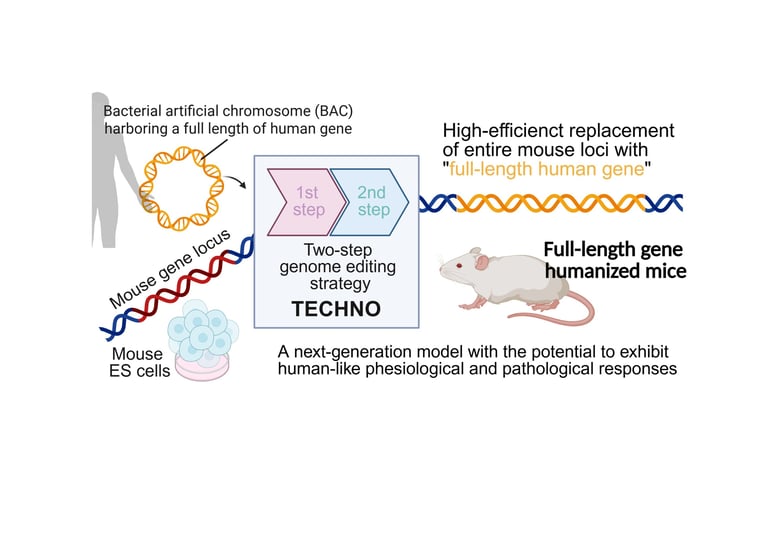
A scalable two-step platform, TECHNO, enables full-length gene humanization by replacing entire mouse loci with their human counterparts using CRISPR/Cas9-assisted editing and BAC-based delivery of large human genomic regions.
The process begins by removing the target locus and inserting human homology arms with a neomycin cassette, then a second step delivers a full-length human genomic fragment via BAC along with a blasticidin cassette to achieve a complete knock-in.
Compared with BAC transgenic approaches, the TECHNO method offers broad applicability to roughly 93% of human genes within BAC limits and notes potential neighboring gene expression effects due to large-scale genomic humanization.
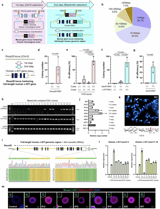
Replacement of the mouse Apobec3 locus with the human APOBEC3 cluster (>200 kb) yielded 10.6–15.2% second-step efficiency across multiple ES cell strains, with expression of all seven human APOBEC3 genes in lung and spleen mirroring human tissue patterns.
In the first step, the mouse locus is excised and replaced with short human homology arms flanking a selectable cassette to create a precise landing site; the second step uses a BAC carrying the full-length human gene and regulatory elements to enable homology-directed integration of fragments over 200 kilobase pairs.
Humanization of the mouse c-Kit locus required longer human homology arms (about 3 kb versus 1 kb) to reach over 10% efficiency, yielding germline transmission with largely preserved KIT function, though some anemia-like features and reduced testis weight appeared in homozygotes.
Applications include humanizing c-Kit, APOBEC3, and CYBB loci, with c-Kit showing human-like alternative splicing and tissue-specific expression, APOBEC3 achieving integration over 200 kb spanning seven genes, and CYBB enabling modeling of chronic granulomatous disease through disease-associated mutations.
Overall, the authors present a scalable ES cell–based full-length gene-humanization platform with robust efficiencies for precise in vivo humanization and disease modeling across multiple loci and haplotypes.
The TECHNO platform accelerates creation of precise, human-relevant mouse models for therapeutic target evaluation, disease-variant validation, and early screening of ineffective drugs, with potential broad impact on translational medicine.
Knock-in efficiencies varied by locus and arm length, with initial Rosa26 experiments showing 60–80% and BAC-mediated second steps achieving 5–15% in some cases, reflecting locus-dependent performance.
Disease modeling was demonstrated by introducing mutations into a humanized CYBB allele (CGD model), producing mice with human-like hCYBB expression and functional ROS defects mirroring CGD.
Knock-ins at the Rosa26 locus for hKIT achieved about 30% efficiency in the second step, with single-copy, correctly recombined integrations confirmed by FISH and sequencing, and confirmed hKIT mRNA and protein expression.
Summary based on 2 sources
Get a daily email with more Science stories
Sources
Phys.org • Jan 17, 2026
Two-step genome editing enables creation of full-length humanized mouse models