Revolutionary Cell Model Predicts Core Processes, Accelerates Cycle Simulation with Dual-GPU Strategy
March 9, 2026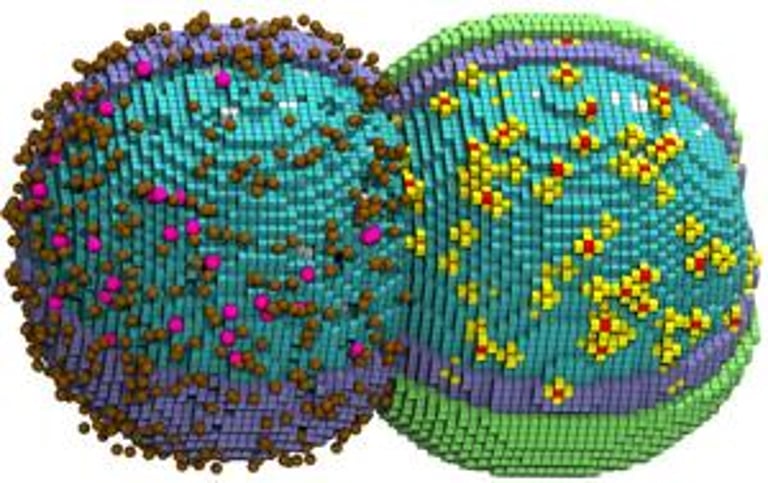
A whole-cell model now enables simultaneous prediction of nucleotide metabolism, DNA replication, and ribosome biogenesis, providing integrated insights into core cellular processes.
The model achieves accurate timing of the cell cycle by averaging molecular dynamics rather than simulating every atom, acknowledging a limitation but highlighting strong predictive power.
This work is part of the NSF Science and Technology Center for Quantitative Cell Biology at the University of Illinois, aiming to advance computer vision approaches to fundamental life processes.
Researchers note the challenge of simulating moving, 3D cellular components and sometimes render interior elements invisible to reveal processes like chromosome threading.
Co-authors include Angad Mehta and Taekjip Ha, with contributions from Zane Thornburg and Andrew Maytin to refine simulations and address real-time cellular events.
Collaborations with Harvard Medical School and the J. Craig Venter Institute, and use of NSF-supported Delta computing resources at the University of Illinois, supported the project.
The study was published in Cell, with accompanying project details available through qcb.illinois.edu and JCVI resources.
The team integrated experimental data from multiple institutions with extensive computational modeling to map thousands of molecular interactions and reactions.
Interdisciplinary collaboration spanned computational biology, chemistry, physics, and experimental biology, leveraging data from Harvard, Illinois, and NSF facilities.
Advances in multiscale modeling reconcile mechanical and biochemical interactions, enabling exploration of cellular responses to stress or mutation and informing synthetic biology and minimal life research.
A dual-GPU strategy accelerated simulation: one GPU handles chromosome replication, another handles metabolism and ribosome assembly, enabling a full 105-minute cycle in six days of computer time.
The modeling relies on high-performance computing and optimized simulations to manage thousands of molecular interactions, with replication requiring dedicated computational resources.
Summary based on 5 sources
Get a daily email with more Science stories
Sources
EurekAlert! • Mar 9, 2026
Team simulates a living cell that grows and divides
News-Medical • Mar 9, 2026
Scientists simulate complete life cycle of a minimal bacterial cell
GEN - Genetic Engineering and Biotechnology News • Mar 9, 2026
Simulating Life: 4D Whole-Cell Model of a Minimal Bacterium